
The intriguing success of helper components in vector-transmission of plant viruses.
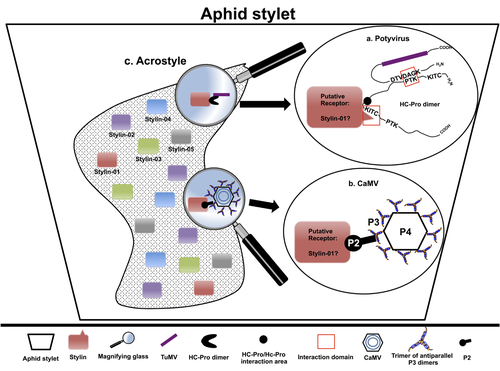
The helper strategy in vector-transmission of plant viruses
Abstract
Recommendation: posted 22 March 2023, validated 23 March 2023
Coustau, C. (2023) The intriguing success of helper components in vector-transmission of plant viruses.. Peer Community in Infections, 100075. https://doi.org/10.24072/pci.infections.100075
Recommendation
Most plant-infecting viruses rely on an animal vector to be transmitted from one sessile host plant to another. A fascinating aspect of virus-vector interactions is the fact that viruses from different clades produce different proteins to bind vector receptors (1). Two major processes are described. In the “capsid strategy”, a motif of the capsid protein is directly binding to the vector receptor. In the “helper strategy”, a non-structural component, the helper component (HC), establishes a bridge between the virus particle and the vector’s receptor.
In this exhaustive review focusing on hemipteran insect vectors, Di Mattia et al. (2) are revisiting the helper strategy in light of recent results. The authors first place the discoveries of the HC strategy in a historical context, suggesting that HC are exclusively found in non-circulative viruses (viruses that only attach to the vector). They present an overview of the nature and modes of action of helper components in the major virus clades of non-circulative viruses (Potyviruses and Caulimoviruses). Authors then detail recent advances, to which they have significantly contributed, showing that the helper strategy also appears widespread in circulative transmission categories (Tenuiviruses, Nanoviruses).
In an extensive perspective section, they raise the question of the evolutionary significance of the existence of HC in numerous unrelated viruses, transmitted by unrelated vectors through different mechanisms. They explore the hypothesis that the helper strategy evolved several times independently in distinct viral clades and for different reasons. In particular, they present several potential benefits of plant virus HC related to virus cooperation, collective transmission and effector-driven infectivity.
As pointed out by both reviewers, this is a very clear and synthetic review. Di Mattia et al. present an exhaustive overview of virus HC-vector molecular interactions and address functionally and evolutionarily important questions. This review should benefit a large audience interested in host-virus interactions and transmission processes.
REFERENCES
(1) Ng JCK, Falk BW (2006) Virus-Vector Interactions Mediating Nonpersistent and Semipersistent Transmission of Plant Viruses. Annual Review of Phytopathology, 44, 183–212. https://doi.org/10.1146/annurev.phyto.44.070505.143325
(2) Di Mattia J, Zeddam J-L, Uzest M, Blanc S (2023) The helper strategy in vector-transmission of plant viruses. Zenodo, ver. 2 peer-reviewed and recommended by Peer Community In Infections. https://doi.org/10.5281/zenodo.7709290
The recommender in charge of the evaluation of the article and the reviewers declared that they have no conflict of interest (as defined in the code of conduct of PCI) with the authors or with the content of the article. The authors declared that they comply with the PCI rule of having no financial conflicts of interest in relation to the content of the article.
This work was funded by the French ANR : REASSORT ANR-20-CE02-0016 and Nanovirus ANR-18-CE92- 0028, as well as by the University of Montpellier MUSE, project MULTIVIR. JDM, SB and MU acknowledge funds from INRAE and JLZ from IRD.
Evaluation round #1
DOI or URL of the preprint: https://doi.org/10.5281/zenodo.7261676
Version of the preprint: 1
Author's Reply, 08 Mar 2023
Dear Recommender
We here attach a pdf file of our point-by-point response to the reviewers' comments
We hope thatt this and the related changes of the manuscript text will be judged acceptable
We thank you very much in advance for your consideration
Sincerely
Stephane Blanc
Decision by Christine Coustau, posted 15 Dec 2022, validated 16 Dec 2022
Dear authors,
We have now received two evaluations of your very interesting review. Both reviewers (and myself) enjoyed reading your review and highlighted its quality.
They also made few suggestions of minor changes that may be taken into account for improving the manuscript. I hope you will find them useful.
Thank you for having submitted your work to PCI Infections.
I look forward to reading the next version of your manuscript.
Sincerely
Christine Coustau"
Reviewed by Jamie Bojko, 15 Dec 2022
Thank you for the opportunity to comment on this review. It was a very interesting read. I’ve listed a few comments below; however, my comments are rather superficial, because the review is well written and builds in a lot of detail.
Broader considerations:
You may choose to broaden the taxonomic detail of the virus groups by including the most recent ICTV publication relative to the virus family you discuss.
In section 3 and 4, perhaps the authors would consider separating the two examples: Potyvirus and Caulimovirus (for example) into two more encompassing sections. This would allow the reader to see the whole process in parallel for two viruses. Providing exemplary figures for this would be even better, but not entirely necessary.
I think that you start your conclusions section a little early. I would consider putting the sections within your conclusion as sections in the main text. Perhaps under a “Hypotheses in HC” heading.
Minor notes:
- You use “thus” and “very” a lot, most often inappropriately in this scientific context. I would remove this from the manuscript to streamline it.
- Your reference system seems to include an additional name in each case. Could this be restricted to the last name of the first author (or two in the case of less than 3 authors) followed by et al. and the year?
- Take care for grammar throughout.
Interesting notes:
- I am interested in your notes on Tenuivirus having two glycoproteins encoded. We also see this on crustacean bunyavirus genomes, but there has been little to no work to determine what their true function is and there are some that are most related to the plant bunyaviruses.
https://doi.org/10.24072/pci.infections.100075.rev11











